Last Updated on 13/05/2026 by Eran Feit
In this guide, we dive deep into the intersection of computer vision and modern agriculture by building a robust system for identifying plant pathologies. This Faster R-CNN PyTorch Tutorial focuses specifically on tomato leaf disease detection, a critical challenge for farmers and agronomists worldwide. By leveraging deep learning, we can automate the diagnosis of common issues like Early Blight and Mosaic Virus, transforming how we monitor crop health at scale.
Developing high-accuracy detection systems provides an essential tool for agricultural researchers and software engineers looking to make a tangible impact on food security. By moving beyond simple classification to precise object detection, readers gain the ability to locate exactly where a disease is manifesting on a single leaf. This technical proficiency is highly sought after in the industry, as it allows for more targeted treatments and significant reductions in chemical usage.
We achieve this by walking through a complete implementation, starting from data preparation with the COCO format to fine-tuning a pre-trained ResNet-50 backbone. This Faster R-CNN PyTorch Tutorial demystifies the process of setting up data loaders, defining model heads, and managing training loops on a GPU. By the end of this article, you will have a functional, end-to-end pipeline that can be adapted for any custom object detection task.
Ultimately, this tutorial serves as a bridge between complex academic papers and practical, deployable code. Whether you are a student exploring neural networks or a developer building a commercial ag-tech solution, the insights provided here offer a clear roadmap. Using the power of PyTorch, we turn raw images into actionable data, ensuring your next computer vision project is built on a foundation of industry-standard best practices.
Why Faster R-CNN is a Game-Changer for Your PyTorch Projects Faster R-CNN remains one of the most influential architectures in the world of computer vision because it successfully solved the bottleneck of “region proposal.” In earlier versions, finding the “interesting” parts of an image was a slow, separate process. However, this Faster R-CNN PyTorch Tutorial utilizes the Region Proposal Network (RPN), which allows the model to “look” at the image and predict where objects might be at the same time it is trying to identify them. This two-stage approach ensures that the model maintains a high degree of spatial accuracy, which is vital when you are trying to spot tiny spots of mold or microscopic pests on a tomato leaf.
The primary target of a Faster R-CNN model is to achieve a balance between localization and classification. While “one-stage” detectors like YOLO are built for extreme speed, Faster R-CNN is often preferred in scientific and medical contexts where precision is more important than real-time frames-per-second. It works by extracting feature maps through a backbone—in our case, ResNet-50—and then refining those areas to produce a final bounding box and a class label. The mathematical foundation of this process involves a multi-task loss function that minimizes both the classification error and the distance between the predicted and actual bounding boxes.
Tomato Leaf Disease Detection: Faster R-CNN PyTorch Tutorial 12 Implementing this in the PyTorch ecosystem is particularly advantageous because of the torchvision library. It provides pre-trained weights that have already “learned” how to see basic shapes, colors, and textures from millions of images. By following a Faster R-CNN PyTorch Tutorial , you are essentially standing on the shoulders of giants, taking a model that already understands the visual world and “fine-tuning” it to become an expert in tomato leaf diseases. This high-level flexibility is why PyTorch is the preferred framework for researchers in the United States and Europe who require both power and ease of use.
Tomato Leaf Disease Detection Deep Learning The Blueprint: How Our Code Identifies Tomato Leaf Diseases What is the core engine driving this detection script? The heart of this script is a fine-tuned Faster R-CNN model with a ResNet-50 backbone, which has been surgically modified to replace its generic classification head with one specifically trained to recognize ten distinct tomato leaf pathologies and healthy tissue.
The code provided in this Faster R-CNN PyTorch Tutorial serves as a complete, end-to-end pipeline designed to take raw agricultural imagery and transform it into a sophisticated diagnostic tool. At its most basic level, the script is architected to solve two simultaneous problems: identifying “where” a leaf is in an image (localization) and “what” is wrong with it (classification). By utilizing the torchvision detection library, we are able to leverage industry-standard architectures that have been proven in competitive computer vision benchmarks, ensuring that our agricultural solution is built on a rock-solid foundation.
To make the data digestible for our neural network, the code implements a custom CocoTransform and get_coco_dataset function. This is a crucial step because deep learning models are picky eaters; they require images to be converted into normalized tensors and annotations to be structured in a very specific dictionary format. By wrapping the standard CocoDetection class, we ensure that our training labels—which include the coordinates for the disease spots—are perfectly aligned with the visual features the model is trying to learn during the training process.
The training logic contained in the train_one_epoch function is where the actual “learning” happens. The script iterates through batches of tomato leaf images, pushing them through the model’s layers to generate predictions. It then calculates the difference between those predictions and the ground-truth annotations using a multi-task loss function. By applying backpropagation through the Stochastic Gradient Descent (SGD) optimizer, the code slowly tweaks the model’s internal weights, effectively teaching it to ignore the background soil or greenhouse equipment and focus strictly on the visual markers of diseases like Late Blight or Spider Mites.
Finally, the testing phase of the script provides the visual proof of the model’s intelligence. By loading the saved model weights (.pth files) into a dedicated evaluation loop, we can run inference on completely new images that the model has never seen before. The code uses matplotlib and OpenCV to draw colored bounding boxes and labels directly onto the leaf images, providing a clear, intuitive output. This bridge from abstract mathematical tensors to a visible “Predicted – Early Blight” label is what makes this Faster R-CNN PyTorch Tutorial an essential resource for anyone looking to apply AI to real-world biological challenges.
Link to the tutorial here
Download the code for the tutorial here or here
Link for Medium users here
Master Computer Vision
Follow my latest tutorials and AI insights on my
Personal Blog .
Beginner Complete CV Bootcamp
Foundation using PyTorch & TensorFlow.
Get Started → Interactive Deep Learning with PyTorch
Hands-on practice in an interactive environment.
Start Learning → Advanced Modern CV: GPT & OpenCV4
Vision GPT and production-ready models.
Go Advanced → Faster R-CNN PyTorch Tutorial Tomato Leaf Disease Detection: Faster R-CNN PyTorch Tutorial Building a high-performance object detection model is one of the most rewarding challenges in computer vision, especially when applied to critical fields like agriculture. In this Faster R-CNN PyTorch Tutorial , we are going to walk through the entire process of detecting tomato leaf diseases using a custom dataset formatted in the COCO standard. This article is designed to bridge the gap between complex deep learning theory and practical, deployable code that solves real-world problems.
Why should you follow along? By the end of this guide, you won’t just have a script; you’ll have a deep understanding of how to manage data transforms, configure a pre-trained ResNet-50 backbone, and run a full training cycle on a GPU. We will use Python and PyTorch to automate the detection of nine different disease classes, helping you build a tool that can eventually be used by farmers to protect their crops.
This guide provides a structured, six-part breakdown of the code. Whether you are a researcher or a software engineer, these steps will show you exactly how to implement a state-of-the-art detector from scratch. Let’s dive into the code and start building your AI crop monitor today.
tomato leaf detection Preparing Your Workbench with the Right Tools Before we can start training our model, we need to ensure our local environment is correctly configured with the necessary libraries. A clean installation ensures that your dependencies don’t conflict, providing a smooth experience as you run the detection scripts. Using a virtual environment is highly recommended to keep your computer vision projects organized and isolated.
The core of our project relies on the PyTorch ecosystem, specifically the torchvision library which contains our Faster R-CNN architecture. We also need utility tools like OpenCV for image manipulation and tqdm to provide those helpful progress bars during the training process. These libraries are the standard toolkit for AI engineers, ensuring your code remains compatible with professional workflows.
If you have an NVIDIA GPU, installing the CUDA-enabled version of PyTorch will significantly speed up your training time. While the code will run on a CPU, the heavy mathematical lifting required by the ResNet-50 backbone is best handled by dedicated hardware. Taking the time to set up your environment now will save you hours of troubleshooting once we get into the deep learning implementation.
Which library is the most critical for this object detection project? The torchvision library is the most essential component because it provides the pre-trained Faster R-CNN model and the specific ResNet-50 weights we need for transfer learning.
- Create conda enviroment conda create -- name Pytorch251 python = 3.12 conda activate Pytorch251 nvcc -- version # Cuda 12.4 conda install pytorch == 2.5 . 1 torchvision == 0.20 . 1 torchaudio == 2.5 . 1 pytorch - cuda = 12.4 - c pytorch - c nvidia pip install opencv - python == 4.11 . 0 . 86 pip install matplotlib == 3.10 . 0 pip install pycocotools == 2.0 . 8 Setting up the right environment is the foundation of any successful computer vision project. This Faster R-CNN PyTorch Tutorial is specifically designed to work with the latest versions of Torchvision, ensuring that your local setup is ready to handle the complex computations required for deep learning. By following these installation steps, you ensure that the rest of the Faster R-CNN PyTorch Tutorial runs smoothly on your hardware.
Getting Your Data Ready for Action Preparing your data is the most important step in any machine learning project because the quality of your input directly determines the quality of your output. In this section, we define the specific disease classes we want to identify, ranging from “Early Blight” to “Yellow Leaf Curl Virus.” By setting up the background class and defining our categories, we provide the framework the model needs to understand what it is looking at.
We also implement a custom transformation class called CocoTransform which converts our raw leaf images into tensors. This is vital because PyTorch models can only process numerical data in tensor format. The code then uses the CocoDetection class to link our image folders with their corresponding JSON annotation files, ensuring every bounding box is correctly matched to its label.
Finally, we organize our data into DataLoader objects. These loaders act as the delivery system for our training process, feeding batches of eight images at a time into the model. This batching strategy is efficient for your GPU’s memory and helps the model generalize better by shuffling the training data during each epoch.
Want the exact dataset so your results match mine? If you want to reproduce the same training flow and compare your results to mine, I can share the dataset structure and what I used in this tutorial. Send me an email and mention the name of this tutorial, so I know what you’re requesting.
🖥️ Email: feitgemel@gmail.com
How do we prepare COCO formatted data for a PyTorch detector? We use the torchvision.datasets.CocoDetection class combined with a custom transform to convert images into tensors and match them with their respective JSON annotations.
### This command imports the main PyTorch library. import torch ### This command imports the computer vision library for PyTorch. import torchvision ### This command imports the utility for loading datasets in batches. from torch . utils . data import DataLoader ### This command imports the Faster R-CNN model class. from torchvision . models . detection import FasterRCNN ### This command imports the specific weights for the ResNet50 backbone. from torchvision . models . detection import FasterRCNN_ResNet50_FPN_Weights ### This command imports the tool to create a new classification head. from torchvision . models . detection . faster_rcnn import FastRCNNPredictor ### This command imports the standard COCO dataset handler. from torchvision . datasets import CocoDetection ### This command imports functional transforms for image manipulation. from torchvision . transforms import functional as F ### This command imports the plotting library for visualization. import matplotlib . pyplot as plt ### This command imports the Python Imaging Library for handling images. from PIL import Image ### This command imports numerical computing tools. import numpy as np ### This command imports progress bar utilities. from tqdm . auto import tqdm ### This command imports the OpenCV library for image processing. import cv2 ### This command imports system utilities for folder management. import os ### This list defines the labels for each tomato leaf disease. classes = [ " Tomato-Leaf " , " Early Blight " , " Healthy " , " Late Blight " , " Leaf Miner " , " Leaf Mold " , " Mosaic Virus " , " Septoria " , " Spider Mites " , " Yellow Leaf Curl Virus " ] ### This variable calculates the total number of classes including the background. num_of_classes = len ( classes ) + 1 # Add backgournd class ### This class defines the transformation applied to each image. class CocoTransform : ### This method handles the conversion when the class is called. def __call__ ( self , image , target ): ### This command converts the image into a PyTorch tensor. image = F . to_tensor ( image ) # Convert PIL image to tensor ### This command returns the processed image and original target. return image , target ### This function initializes the COCO dataset with specific paths. def get_coco_dataset ( img_dir , ann_file ): # params : image folder and the annotation file ### This command returns a CocoDetection object with the defined transform. return CocoDetection ( root = img_dir , annFile = ann_file , transforms = CocoTransform ()) ### This variable loads the training images and annotations. train_dataset = get_coco_dataset ( img_dir = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/train " , ann_file = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/train/_annotations.coco.json " ) ### This variable loads the validation images and annotations. val_dataset = get_coco_dataset ( img_dir = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/valid " , ann_file = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/valid/_annotations.coco.json " ) ### This command creates a data loader for the training set. train_loader = DataLoader ( train_dataset , batch_size = 8 , shuffle =True , collate_fn =lambda x : tuple ( zip ( * x ))) ### This command creates a data loader for the validation set. val_loader = DataLoader ( val_dataset , batch_size = 8 , shuffle =False , collate_fn =lambda x : tuple ( zip ( * x ))) ### This command prints the total number of training images. print ( f "Number of training images : { len ( train_dataset ) } " ) ### This command prints the total number of validation images. print ( f "Number of validation images : { len ( val_dataset ) } " ) Seeing Through the Eyes of Your Dataset Before we start the heavy lifting of training, it is crucial to visualize our data to ensure our labels are accurate. In this part of the code, we grab the very first image from our training set and prepare it for display using OpenCV. Since PyTorch and OpenCV handle image data in different formats, we perform a “permutation” and “normalization” to make the leaf image look right on your screen.
The loop inside this section iterates through every annotation associated with that image. It extracts the bounding box coordinates—represented in COCO’s [x, y, width, height] format—and converts them into the pixel coordinates OpenCV needs to draw rectangles. Seeing these green boxes appear on your screen is your first confirmation that the data pipeline is working correctly.
This visual check is a developer’s best friend. It allows you to verify that the “Late Blight” label is actually hovering over a diseased spot and not a random piece of dirt. By manually inspecting these samples, we ensure that the model has a clear and accurate set of instructions to learn from during the training epochs.
Why do we need to convert PyTorch tensors back to NumPy for visualization? OpenCV and Matplotlib expect images in a HWC (Height, Width, Channels) format with values from 0-255, while PyTorch uses CHW (Channels, Height, Width) format with normalized values.
### This command imports OpenCV for visualization. import cv2 ### This command imports NumPy for array manipulation. import numpy as np ### This command retrieves the first sample from the dataset. image , target = train_dataset [ 0 ] # First sample ### This command reorders the tensor dimensions for OpenCV. image_np = image . permute ( 1 , 2 , 0 ). numpy () ### This command scales the pixel values back to the 0-255 range. image_np = ( image_np * 255 ). astype ( np . uint8 ) ### This command converts the color space from RGB to BGR. image_np = cv2 . cvtColor ( image_np , cv2 . COLOR_RGB2BGR ) ### This loop iterates through every annotated object in the image. for obj in target : # Traget is a list ### This command extracts the bounding box coordinates. bbox = obj [ " bbox " ] # COCO format : [x , y, width, height] ### This command converts the coordinates into integer values. x , y , w , h = map ( int , bbox ) # Convert to integers ### This variable calculates the bottom-right corner of the box. x_max , y_max = x + w , y + h ### This command draws a green rectangle on the image. cv2 . rectangle ( image_np , ( x , y ) , ( x_max , y_max ) , ( 0 , 255 , 0 ) , 2 ) ### This command opens a window to show the annotated image. cv2 . imshow ( " First train image with bounding boxes " , image_np ) ### This command waits for a key press before closing the window. cv2 . waitKey ( 0 ) ### This command closes all open OpenCV windows. cv2 . destroyAllWindows () Choosing the Right Neural Architecture To detect complex patterns like leaf mold or spider mites, we need a model that is both deep and flexible. In this section, we initialize the Faster R-CNN architecture with a ResNet-50-FPN backbone. Faster R-CNN is a two-stage detector, which means it first proposes regions where objects might be and then classifies those regions, making it incredibly accurate for high-detail tasks.
We don’t build this model from scratch; instead, we load pre-trained weights that have already learned to see basic shapes on millions of images. However, since the original model was designed for 80 generic classes (like dogs or cars), we replace the “box predictor” head with a custom one that matches our eleven tomato-specific classes. This process, known as Transfer Learning, allows us to get professional results with a much smaller dataset.
Once the model is built, we move it to the GPU (CUDA) for lightning-fast calculations. We also set up a Stochastic Gradient Descent (SGD) optimizer and a learning rate scheduler. The scheduler is particularly clever; it automatically reduces the learning rate as training progresses, allowing the model to “settle in” and find the absolute best mathematical solution for detection.
Selecting the ResNet-50-FPN backbone is a strategic choice for this Faster R-CNN PyTorch Tutorial . While there are many architectures available, this specific combination offers an excellent balance between training stability and detection accuracy. As we move forward in the Faster R-CNN PyTorch Tutorial , you will see how this backbone allows the model to extract high-level features from tomato leaves with incredible efficiency.
How do we modify a pre-trained model for a specific number of classes? We extract the number of input features from the existing classifier and then replace the box_predictor with a new FastRCNNPredictor that uses our custom class count.
### This function builds the model and replaces the classification head. def get_model ( num_classes ): ### This command loads the pre-trained Faster R-CNN architecture. model = torchvision . models . detection . fasterrcnn_resnet50_fpn ( weights = FasterRCNN_ResNet50_FPN_Weights . DEFAULT ) ### This variable stores the number of features in the original classifier. in_featurs = model . roi_heads . box_predictor . cls_score . in_features ### This command overwrites the original head with our custom predictor. model . roi_heads . box_predictor = FastRCNNPredictor ( in_featurs , num_classes ) ### This command returns the modified model. return model ### This variable initializes our custom model. model = get_model ( num_of_classes ) ### This variable selects the GPU if available or the CPU otherwise. device = torch . device ( ' cuda ' ) if torch . cuda . is_available () else torch . device ( ' cpu ' ) ### This command transfers the model parameters to the selected device. model . to ( device ) ### This list identifies the model parameters that need to be trained. params = [ p for p in model . parameters () if p . requires_grad ] ### This command initializes the optimizer with specific learning settings. optimizer = torch . optim . SGD ( params , lr = 0.005 , momentum = 0.9 , weight_decay = 0.0005 ) ### This command creates a scheduler to decrease the learning rate over time. lr_scheduler = torch . optim . lr_scheduler . StepLR ( optimizer , step_size = 3 , gamma = 0.1 ) Tomato Leaf Disease Detection: Faster R-CNN PyTorch Tutorial Building a high-performance object detection model is one of the most rewarding challenges in computer vision, especially when applied to critical fields like agriculture. In this Faster R-CNN PyTorch Tutorial , we are going to walk through the entire process of detecting tomato leaf diseases using a custom dataset formatted in the COCO standard. This article is designed to bridge the gap between complex deep learning theory and practical, deployable code that solves real-world problems.
Why should you follow along? By the end of this guide, you won’t just have a script; you’ll have a deep understanding of how to manage data transforms, configure a pre-trained ResNet-50 backbone, and run a full training cycle on a GPU. We will use Python and PyTorch to automate the detection of nine different disease classes, helping you build a tool that can eventually be used by farmers to protect their crops.
This guide provides a structured, six-part breakdown of the code. Whether you are a researcher or a software engineer, these steps will show you exactly how to implement a state-of-the-art detector from scratch. Let’s dive into the code and start building your AI crop monitor today.
Getting Your Data Ready for Action Preparing your data is the most important step in any machine learning project because the quality of your input directly determines the quality of your output. In this section, we define the specific disease classes we want to identify, ranging from “Early Blight” to “Yellow Leaf Curl Virus.” By setting up the background class and defining our categories, we provide the framework the model needs to understand what it is looking at.
We also implement a custom transformation class called CocoTransform which converts our raw leaf images into tensors. This is vital because PyTorch models can only process numerical data in tensor format. The code then uses the CocoDetection class to link our image folders with their corresponding JSON annotation files, ensuring every bounding box is correctly matched to its label.
Finally, we organize our data into DataLoader objects. These loaders act as the delivery system for our training process, feeding batches of eight images at a time into the model. This batching strategy is efficient for your GPU’s memory and helps the model generalize better by shuffling the training data during each epoch.
Want the exact dataset so your results match mine? If you want to reproduce the same training flow and compare your results to mine, I can share the dataset structure and what I used in this tutorial. Send me an email and mention the name of this tutorial, so I know what you’re requesting.
🖥️ Email: feitgemel@gmail.com
How do we prepare COCO formatted data for a PyTorch detector? We use the torchvision.datasets.CocoDetection class combined with a custom transform to convert images into tensors and match them with their respective JSON annotations.
Python
### This command imports the main PyTorch library. import torch ### This command imports the computer vision library for PyTorch. import torchvision ### This command imports the utility for loading datasets in batches. from torch . utils . data import DataLoader ### This command imports the Faster R-CNN model class. from torchvision . models . detection import FasterRCNN ### This command imports the specific weights for the ResNet50 backbone. from torchvision . models . detection import FasterRCNN_ResNet50_FPN_Weights ### This command imports the tool to create a new classification head. from torchvision . models . detection . faster_rcnn import FastRCNNPredictor ### This command imports the standard COCO dataset handler. from torchvision . datasets import CocoDetection ### This command imports functional transforms for image manipulation. from torchvision . transforms import functional as F ### This command imports the plotting library for visualization. import matplotlib . pyplot as plt ### This command imports the Python Imaging Library for handling images. from PIL import Image ### This command imports numerical computing tools. import numpy as np ### This command imports progress bar utilities. from tqdm . auto import tqdm ### This command imports the OpenCV library for image processing. import cv2 ### This command imports system utilities for folder management. import os ### This list defines the labels for each tomato leaf disease. classes = [ " Tomato-Leaf " , " Early Blight " , " Healthy " , " Late Blight " , " Leaf Miner " , " Leaf Mold " , " Mosaic Virus " , " Septoria " , " Spider Mites " , " Yellow Leaf Curl Virus " ] ### This variable calculates the total number of classes including the background. num_of_classes = len ( classes ) + 1 # Add backgournd class ### This class defines the transformation applied to each image. class CocoTransform : ### This method handles the conversion when the class is called. def __call__ ( self , image , target ): ### This command converts the image into a PyTorch tensor. image = F . to_tensor ( image ) # Convert PIL image to tensor ### This command returns the processed image and original target. return image , target ### This function initializes the COCO dataset with specific paths. def get_coco_dataset ( img_dir , ann_file ): # params : image folder and the annotation file ### This command returns a CocoDetection object with the defined transform. return CocoDetection ( root = img_dir , annFile = ann_file , transforms = CocoTransform ()) ### This variable loads the training images and annotations. train_dataset = get_coco_dataset ( img_dir = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/train " , ann_file = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/train/_annotations.coco.json " ) ### This variable loads the validation images and annotations. val_dataset = get_coco_dataset ( img_dir = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/valid " , ann_file = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/valid/_annotations.coco.json " ) ### This command creates a data loader for the training set. train_loader = DataLoader ( train_dataset , batch_size = 8 , shuffle =True , collate_fn =lambda x : tuple ( zip ( * x ))) ### This command creates a data loader for the validation set. val_loader = DataLoader ( val_dataset , batch_size = 8 , shuffle =False , collate_fn =lambda x : tuple ( zip ( * x ))) ### This command prints the total number of training images. print ( f "Number of training images : { len ( train_dataset ) } " ) ### This command prints the total number of validation images. print ( f "Number of validation images : { len ( val_dataset ) } " ) Seeing Through the Eyes of Your Dataset Before we start the heavy lifting of training, it is crucial to visualize our data to ensure our labels are accurate. In this part of the code, we grab the very first image from our training set and prepare it for display using OpenCV. Since PyTorch and OpenCV handle image data in different formats, we perform a “permutation” and “normalization” to make the leaf image look right on your screen.
The loop inside this section iterates through every annotation associated with that image. It extracts the bounding box coordinates—represented in COCO’s [x, y, width, height] format—and converts them into the pixel coordinates OpenCV needs to draw rectangles. Seeing these green boxes appear on your screen is your first confirmation that the data pipeline is working correctly.
This visual check is a developer’s best friend. It allows you to verify that the “Late Blight” label is actually hovering over a diseased spot and not a random piece of dirt. By manually inspecting these samples, we ensure that the model has a clear and accurate set of instructions to learn from during the training epochs.
Why do we need to convert PyTorch tensors back to NumPy for visualization? OpenCV and Matplotlib expect images in a HWC (Height, Width, Channels) format with values from 0-255, while PyTorch uses CHW (Channels, Height, Width) format with normalized values.
Python
### This command imports OpenCV for visualization. import cv2 ### This command imports NumPy for array manipulation. import numpy as np ### This command retrieves the first sample from the dataset. image , target = train_dataset [ 0 ] # First sample ### This command reorders the tensor dimensions for OpenCV. image_np = image . permute ( 1 , 2 , 0 ). numpy () ### This command scales the pixel values back to the 0-255 range. image_np = ( image_np * 255 ). astype ( np . uint8 ) ### This command converts the color space from RGB to BGR. image_np = cv2 . cvtColor ( image_np , cv2 . COLOR_RGB2BGR ) ### This loop iterates through every annotated object in the image. for obj in target : # Traget is a list ### This command extracts the bounding box coordinates. bbox = obj [ " bbox " ] # COCO format : [x , y, width, height] ### This command converts the coordinates into integer values. x , y , w , h = map ( int , bbox ) # Convert to integers ### This variable calculates the bottom-right corner of the box. x_max , y_max = x + w , y + h ### This command draws a green rectangle on the image. cv2 . rectangle ( image_np , ( x , y ) , ( x_max , y_max ) , ( 0 , 255 , 0 ) , 2 ) ### This command opens a window to show the annotated image. cv2 . imshow ( " First train image with bounding boxes " , image_np ) ### This command waits for a key press before closing the window. cv2 . waitKey ( 0 ) ### This command closes all open OpenCV windows. cv2 . destroyAllWindows () HTML
< div class= " ef-related " > < p class= " ef-heading " > MASTER OBJECT DETECTION </ p > < ul class= " ef-links " > < li > < a href = " https://eranfeit.net/accuracy-vs-speed-comparing-faster-r-cnn-and-ssd-in-pytorch/ " target = " _blank " rel = " noopener " > Accuracy vs . Speed : Comparing Faster R - CNN and SSD in PyTorch . </ a > < p > Learn how Faster R - CNN compares to other popular architectures in terms of precision and performance . </ p > </ li > < li > < a href = " https://eranfeit.net/how-to-train-convnext-in-pytorch-on-a-custom-dataset/ " target = " _blank " rel = " noopener " > How to Train ConvNeXt in PyTorch on a Custom Dataset </ a > < p > Discover how to handle custom image datasets in PyTorch using modern classification backbones . </ p > </ li > </ ul > </ div > Choosing the Right Neural Architecture To detect complex patterns like leaf mold or spider mites, we need a model that is both deep and flexible. In this section, we initialize the Faster R-CNN architecture with a ResNet-50-FPN backbone. Faster R-CNN is a two-stage detector, which means it first proposes regions where objects might be and then classifies those regions, making it incredibly accurate for high-detail tasks.
We don’t build this model from scratch; instead, we load pre-trained weights that have already learned to see basic shapes on millions of images. However, since the original model was designed for 80 generic classes (like dogs or cars), we replace the “box predictor” head with a custom one that matches our eleven tomato-specific classes. This process, known as Transfer Learning, allows us to get professional results with a much smaller dataset.
Once the model is built, we move it to the GPU (CUDA) for lightning-fast calculations. We also set up a Stochastic Gradient Descent (SGD) optimizer and a learning rate scheduler. The scheduler is particularly clever; it automatically reduces the learning rate as training progresses, allowing the model to “settle in” and find the absolute best mathematical solution for detection.
How do we modify a pre-trained model for a specific number of classes? We extract the number of input features from the existing classifier and then replace the box_predictor with a new FastRCNNPredictor that uses our custom class count.
Python
### This function builds the model and replaces the classification head. def get_model ( num_classes ): ### This command loads the pre-trained Faster R-CNN architecture. model = torchvision . models . detection . fasterrcnn_resnet50_fpn ( weights = FasterRCNN_ResNet50_FPN_Weights . DEFAULT ) ### This variable stores the number of features in the original classifier. in_featurs = model . roi_heads . box_predictor . cls_score . in_features ### This command overwrites the original head with our custom predictor. model . roi_heads . box_predictor = FastRCNNPredictor ( in_featurs , num_classes ) ### This command returns the modified model. return model ### This variable initializes our custom model. model = get_model ( num_of_classes ) ### This variable selects the GPU if available or the CPU otherwise. device = torch . device ( ' cuda ' ) if torch . cuda . is_available () else torch . device ( ' cpu ' ) ### This command transfers the model parameters to the selected device. model . to ( device ) ### This list identifies the model parameters that need to be trained. params = [ p for p in model . parameters () if p . requires_grad ] ### This command initializes the optimizer with specific learning settings. optimizer = torch . optim . SGD ( params , lr = 0.005 , momentum = 0.9 , weight_decay = 0.0005 ) ### This command creates a scheduler to decrease the learning rate over time. lr_scheduler = torch . optim . lr_scheduler . StepLR ( optimizer , step_size = 3 , gamma = 0.1 ) Teaching the Model to Spot the Difference The heart of our Faster R-CNN PyTorch Tutorial is the training function. This block of code is responsible for taking the images and their labels and teaching the model to minimize its mistakes. We use the model.train() mode, which tells PyTorch that we are currently in learning mode and want to calculate the gradients of the loss function.
Inside the loop, we perform a critical cleanup step: we filter out any images that don’t have valid bounding boxes. This prevents the model from getting confused by empty data. For every valid leaf, we convert the bounding box and class ID into tensors and ship them to the GPU. This ensures that the entire calculation—from image features to loss values—happens in high-speed video memory.
The core of the process is the losses.backward() and optimizer.step() commands. The first one calculates how much each neural connection contributed to the error, and the second one slightly adjusts those connections to perform better next time. We use the tqdm progress bar to keep track of the average loss, giving us real-time feedback on how well the model is learning to identify tomato diseases.
What happens inside the training loop during each batch? The model generates predictions, calculates the loss between those predictions and the ground truth, and then uses backpropagation to update its weights via the optimizer.
### This function manages the training process for one full pass of data. def train_one_epoch ( model , optimizer , data_loader , device , epoch ): ### This command puts the model into training mode. model . train () ### This variable tracks the total loss across the epoch. total_loss = 0 ### This variable counts the number of batches processed. batch_count = 0 ### This variable counts images that were skipped due to errors. skipped_batches = 0 ### This variable tracks the total number of bounding boxes seen. total_boxes = 0 ### This command creates a visual progress bar for the console. progress_bar = tqdm ( data_loader , desc = f "Epoch { epoch + 1 } / { num_epochs } " , leave =True ) ### This loop iterates through the images and targets in the loader. for images , targets in progress_bar : ### This command moves the current batch of images to the GPU. images = [ img . to ( device ) for img in images ] ### This list stores the filtered targets for training. processed_targets = [] ### This list stores the images that have valid targets. valid_images = [] ### This variable counts boxes in the current batch. batch_boxes = 0 ### This loop processes each individual image and its annotations. for i , target in enumerate ( targets ): ### This list stores valid bounding boxes for the image. boxes = [] ### This list stores the class labels for the image. labels = [] ### This loop parses each object in the target annotation. for obj in target : ### This variable extracts the raw box coordinates. bbox = obj [ " bbox " ] # Format : [x, y, width , height] ### This command unpacks the coordinates for conversion. x , y , w , h , = bbox ### This condition ensures we only use boxes with a real area. if w > 0 and h > 0 : ### This command appends the box in min-max format. boxes . append ([ x , y , x + w , y + h ]) # Convert to [x_min, y_min , x_max, y_max] ### This command appends the category ID for the box. labels . append ( obj [ " category_id " ]) ### This condition checks if any valid boxes were found. if boxes : ### This dictionary stores the tensors required by Faster R-CNN. processed_target = { " boxes " : torch . tensor ( boxes , dtype = torch . float32 ). to ( device ), " labels " : torch . tensor ( labels , dtype = torch . int64 ). to ( device ), } ### This command adds the target to the batch list. processed_targets . append ( processed_target ) ### This command adds the image to the valid batch list. valid_images . append ( images [ i ]) ### This command increments the total box count. batch_boxes += len ( boxes ) else : ### This command updates the progress bar with skip info. progress_bar . set_postfix_str ( f "Skipping image { i } (no valid boxes)" ) ### This condition skips the iteration if the batch is empty. if not processed_targets : ### This command increments the skipped batch counter. skipped_batches += 1 ### This command updates the progress bar status. progress_bar . set_postfix_str ( " Skipping batch - No valid targets " ) ### This command reassigns images to the filtered list. images = valid_images ### This command updates the global box count. total_boxes += batch_boxes ### This command calculates the model loss for the batch. loss_dict = model ( images , processed_targets ) ### This command sums the various loss components. losses = sum ( loss for loss in loss_dict . values ()) ### This command clears previous gradients before backpropagation. optimizer . zero_grad () ### This command calculates the gradient of the loss. losses . backward () ### This command updates the model weights. optimizer . step () ### This command adds the batch loss to the total. total_loss += losses . item () ### This command increments the batch counter. batch_count += 1 ### This command updates the progress bar with live metrics. progress_bar . set_postfix ({ ' loss ' : f ' { losses . item () :.4f } ' , ' avg_loss ' : f ' { total_loss / batch_count :.4f } ' , ' boxes ' : total_boxes , ' skipped ' : skipped_batches }) ### This variable calculates the final average loss for the epoch. avg_loss = total_loss / max ( batch_count , 1 ) ### This command prints a detailed summary of the epoch results. tqdm . write ( f " \n [Epoch { epoch + 1 } ] " f "Processed: { batch_count } / { len ( data_loader ) } batches " f "( { skipped_batches } skipped) | " f "Total boxes: { total_boxes } | " f "Average loss: { avg_loss :.4f } " ) ### This command returns the epoch loss for tracking. return avg_loss Crossing the Finish Line with Saved Weights Now that we have defined our training logic, it’s time to run the engine. In this part of the script, we execute the training loop for five epochs. Five might seem like a small number, but thanks to our pre-trained ResNet-50 weights, the model quickly learns the high-level features of tomato diseases without needing thousands of iterations.
After every single epoch, we save the model’s “state dictionary” to your hard drive. This is an essential safety measure. If your computer crashes or the power goes out, you won’t lose your progress. Moreover, these saved .pth files allow you to load the model later for inference without ever having to re-train it, making your AI tool portable and ready for deployment.
As the training runs, the code updates the learning rate using our scheduler and prints a summary of the loss progression. Watching the “average loss” number go down is the ultimate proof that your AI is getting smarter. By the end of the fifth epoch, you’ll have a set of weights that are finely tuned to distinguish between healthy leaves and those suffering from Mosaic Virus or Late Blight.
Why is it important to save the model after every epoch? Saving at each epoch allows you to pick the “best” version of the model and provides a checkpoint in case the training process is interrupted.
### This list stores the loss values for each completed epoch. losses = [] ### This variable sets the total number of training cycles. num_epochs = 5 ### This command initializes the master progress bar for the session. epoch_bar = tqdm ( range ( num_epochs ), desc = " Training Progress " , position = 0 ) ### This command prints the initial configuration for the user. print ( " ======== Start Training ======== " ) print ( f "Model: Faster R-CNN with Resnet-50 backbone " ) print ( f "Number of classes : { num_of_classes } " ) print ( f "Training images: { len ( train_dataset ) } " ) print ( f "Validation images: { len ( val_dataset ) } " ) print ( f "Number of epochs : { num_epochs } " ) print ( " ================================== " ) ### This loop runs the training process for the specified number of epochs. for epoch in epoch_bar : ### This command executes the training function and gets the loss. avg_loss = train_one_epoch ( model , optimizer , train_loader , device , epoch ) ### This command appends the loss to the history list. losses . append ( avg_loss ) ### This command tells the scheduler to update the learning rate. lr_scheduler . step () ### This variable defines the folder path for saving models. model_save_dir = " d:/temp/models/Tomato-Leaf-Disease " ### This command creates the save directory if it doesn't exist. os . makedirs ( model_save_dir , exist_ok =True ) ### This variable defines the full path for the specific epoch file. model_path = f "d:/temp/models/Tomato-Leaf-Disease/Faster_rcnn_resnet50_epoch_ { epoch + 1 } .pth" ### This command saves the model weights to the disk. torch . save ( model . state_dict () , model_path ) ### This command updates the progress bar with the latest stats. epoch_bar . set_postfix ({ ' current_loss ' : f ' { avg_loss :.4f } ' , ' lr ' : f ' { optimizer . param_groups [ 0 ][ " lr " ] :.6f } ' }) ### This command prints the save confirmation to the console. tqdm . write ( f "Model saved: { model_path } " ) ### This command prints the final summary after all epochs are done. print ( " \n ================== Training Complete =================== " ) ### This command displays the loss history across the entire session. print ( f "Loss progression across epochs: { [ f ' { loss :.4f } ' for loss in losses ] } " ) tomato leaf detection results Proving It Works with Fresh Predictions After the training is complete, the most critical step is validation. This part of the Faster R-CNN PyTorch Tutorial focuses on real-world application, taking the weights we just trained and applying them to unseen data. It is here that the true power of the Faster R-CNN PyTorch Tutorial becomes visible, as we transition from abstract loss numbers to concrete, visual bounding boxes.
The final step is the most exciting: testing our AI on images it has never seen before. In this part of the tutorial, we load our best saved weights into the model and switch it to eval() mode. This is a critical switch; it turns off certain behaviors like “Dropout” that are only meant for training, ensuring the model gives consistent and stable predictions.
We select six random images from the test folder and run them through our “Inference” pipeline. The script uses a confidence threshold of 0.5 , meaning it only shows boxes if it’s more than 50% sure it has found a disease. For every detection, the model returns a bounding box, a class label, and a confidence score, which we then draw in red over the image.
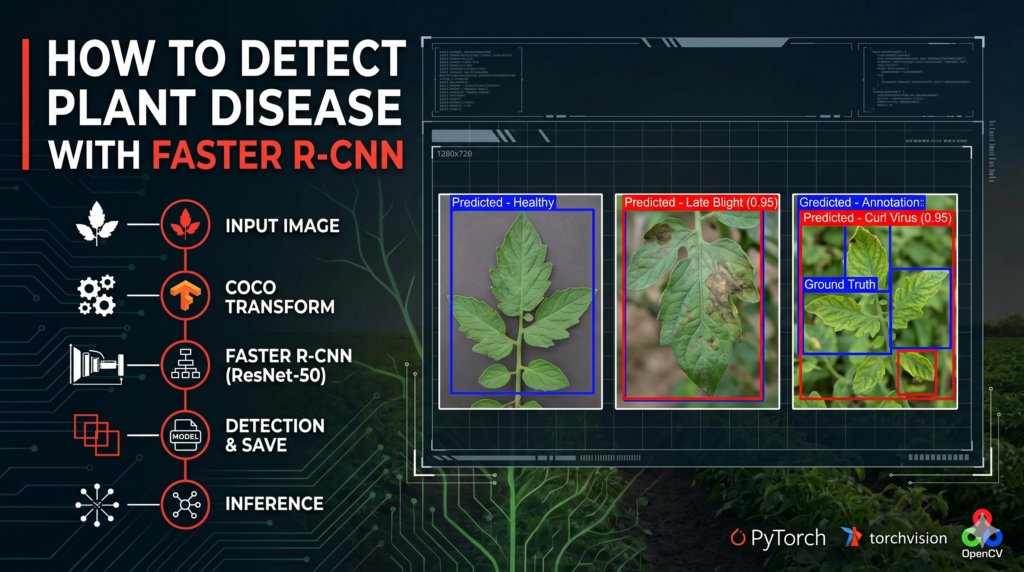
To provide a fair comparison, we also draw the “Ground Truth” (the correct answers) in blue. When the red and blue boxes overlap perfectly, you know your Faster R-CNN PyTorch Tutorial has been a success! This visual dashboard is the perfect way to demonstrate the power of your model to clients or researchers, proving that AI can indeed help protect our food supply.
How do we visualize the final performance of the model? We run the model on test images, draw the predicted boxes in red, and overlay the actual ground-truth boxes in blue using Matplotlib.
### This command puts the model into evaluation mode for testing. model . eval () ### This function prepares a single image for model inference. def prepare_image ( image_path ): ### This command opens the image and ensures it is in RGB format. image = Image . open ( image_path ). convert ( " RGB " ) ### This command converts the image to a tensor and adds a batch dimension. image_tensor = F . to_tensor ( image ). unsqueeze ( 0 ) ### This command moves the tensor to the active device. return image_tensor . to ( device ) ### This variable defines the path to the testing data. test_path = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/test " ### This variable defines the path to the testing annotations. annotation_path = " D:/Data-Sets-Object-Detection/Tomato Leaf Disease.v63i.coco/test/_annotations.coco.json " ### This command imports JSON for parsing annotation files. import json ### This command imports random for image selection. import random ### This block opens and reads the COCO annotation file. with open ( annotation_path , " r " ) as f : coco_annotations = json . load ( f ) ### This dictionary maps image IDs to their actual filenames. image_id_to_filename = { img [ ' id ' ]: img [ ' file_name ' ] for img in coco_annotations [ ' images ' ]} ### This dictionary maps filenames back to their unique IDs. image_filename_to_id = { filename : img_id for img_id , filename in image_id_to_filename . items ()} ### This list identifies all available image files in the test folder. all_images = [ img for img in os . listdir ( test_path ) if img . endswith (( " .jpg " , " .png " ))] ### This command selects six images at random for our dashboard. random_images = random . sample ( all_images , 6 ) ### This dictionary creates a mapping from IDs to class names. COCO_CLASSES = { i + 1 : cls for i , cls in enumerate ( classes ) } ### This command manually adds the background class at index zero. COCO_CLASSES [ 0 ] = " Background " ### This function retrieves the correct labels for a given image. def get_ground_truth_annotations ( filename ): ### This variable looks up the ID for the provided filename. image_id = image_filename_to_id . get ( filename ) ### This list will store all true annotations for the image. ground_truth_annotations = [] ### This loop searches the master list for matching image IDs. for annotation in coco_annotations [ ' annotations ' ]: if annotation [ ' image_id ' ] == image_id : ### This command unpacks the COCO box format. x , y , width , height = annotation [ ' bbox ' ] ### This command converts the box to min-max format. ground_truth_annotations . append ({ ' bbox ' : [ x , y , x + width , y + height ], ' category_id ' : annotation [ ' category_id ' ] }) ### This command returns the list of correct boxes. return ground_truth_annotations ### This function handles the drawing of boxes on the plot. def draw_boxes ( ax , image , prediction , ground_truth ): ### This command displays the base image on the axis. ax . imshow ( image ) ### This loop draws each blue ground-truth box. for gt_box in ground_truth : x_min , y_min , x_max , y_max = gt_box [ ' bbox ' ] class_name = COCO_CLASSES . get ( gt_box [ ' category_id ' ], " Unknown " ) text = f "True - { class_name } " ax . add_patch ( plt . Rectangle (( x_min , y_min ), x_max - x_min , y_max - y_min , linewidth = 2 , edgecolor = ' b ' , facecolor = ' none ' )) ax . text ( x_min , y_min - 20 , text , color = ' blue ' , fontsize = 10 , bbox = dict ( facecolor = ' white ' , alpha = 0.8 , edgecolor = ' none ' , pad = 2 )) ### This block extracts the predicted coordinates and scores. boxes = prediction [ 0 ][ ' boxes ' ]. cpu (). numpy () labels = prediction [ 0 ][ ' labels ' ]. cpu (). numpy () scores = prediction [ 0 ][ ' scores ' ]. cpu (). numpy () ### This variable sets the minimum confidence for a valid detection. threshold = 0.5 ### This loop draws each red predicted box if the score is high enough. for box , label , score in zip ( boxes , labels , scores ): if score > threshold : x_min , y_min , x_max , y_max = box class_name = COCO_CLASSES . get ( label , " Unknown " ) text = f "Predicted - { class_name } ( { score :.2f } )" ax . add_patch ( plt . Rectangle (( x_min , y_min ), x_max - x_min , y_max - y_min , linewidth = 2 , edgecolor = ' r ' , facecolor = ' none ' )) ax . text ( x_min , y_min , text , color = ' red ' , fontsize = 10 , bbox = dict ( facecolor = ' white ' , alpha = 0.8 , edgecolor = ' none ' , pad = 2 )) ### This command hides the axis lines for a cleaner look. ax . axis ( ' off ' ) ### This command creates a 2x3 grid of subplots for visualization. fig , axes = plt . subplots ( 2 , 3 , figsize = ( 20 , 14 ) ) ### This loop iterates through the random images and generates results. for ax , img_name in zip ( axes . flatten () , random_images ): img_path = os . path . join ( test_path , img_name ) image_tensor = prepare_image ( img_path ) ground_truth = get_ground_truth_annotations ( img_name ) ### This block runs the image through the model without gradients. with torch . no_grad (): prediction = model ( image_tensor ) image = Image . open ( img_path ) draw_boxes ( ax , image , prediction , ground_truth ) ### This command optimizes the layout of the final plot. plt . tight_layout () ### This command displays the final results grid. plt . show () FAQ Why is Faster R-CNN chosen for this specific task? Faster R-CNN is a two-stage detector that excels at precision, which is vital for identifying small, subtle disease spots on tomato leaves that one-stage detectors might miss.
What role do the ResNet-50-FPN weights play? These pre-trained weights allow the model to start with a deep understanding of shapes and textures, reducing the amount of data and time needed to train the detector from scratch.
Why should I format my dataset in COCO JSON? COCO is an industry-standard format that integrates seamlessly with PyTorch’s `torchvision` utilities, making data loading and augmentation much more reliable and organized.
Is 5 epochs enough for a custom leaf disease detector? Yes, because we are using transfer learning. Fine-tuning a pre-trained model allows it to reach high accuracy for specialized tasks like agriculture much faster than a cold start.
How does the confidence threshold affect the visual output? The threshold filters out low-probability detections; setting it to 0.5 ensures that only predictions where the model is at least 50% sure are drawn as bounding boxes.
Do I need an NVIDIA GPU to follow this tutorial? While a GPU is highly recommended for speed, the provided code automatically detects your hardware and will fall back to the CPU if no CUDA-compatible device is found.
What is the benefit of saving the model after every epoch? Saving `.pth` files creates checkpoints, allowing you to recover from a system crash or compare different versions of the model to find the one with the lowest loss.
Why does the code skip images with no valid bounding boxes? Empty annotations can confuse the model’s loss calculation. Filtering them out ensures the optimization process stays focused on real disease features rather than background noise.
What is the difference between training and evaluation mode? Evaluation mode (`model.eval()`) tells PyTorch to disable training-only features like Dropout, ensuring your predictions are consistent and stable for testing.
How can I use these results for my own custom detection data? You can adapt this pipeline to any project by simply updating the image directory paths and the `classes` list to match your specific dataset labels.
Conclusion: Bridging AI and Agriculture Completing this Faster R-CNN PyTorch Tutorial is a significant milestone in your computer vision journey. Throughout this guide, we have moved from raw data loading to the sophisticated task of fine-tuning a two-stage object detector. By successfully implementing the Faster R-CNN architecture, you have built a tool capable of recognizing subtle biological patterns that are often difficult even for human eyes to spot consistently. This project demonstrates that deep learning isn’t just for academic research—it’s a practical solution for securing global food supplies.
The power of this workflow lies in its flexibility. While we focused on tomato leaves, the same code and logic can be applied to any object detection task, whether you are tracking athletes, monitoring wildlife, or detecting defects in industrial manufacturing. The use of the COCO format and the ResNet-50 backbone ensures that your project follows industry-standard best practices, making your code easier to maintain and scale. As you continue to refine your model, consider experimenting with more epochs or larger datasets to push the boundaries of your AI’s accuracy.
Masterfully identifying crop diseases is just the beginning of what you can achieve with these skills. I hope this Faster R-CNN PyTorch Tutorial has provided you with a clear, actionable roadmap for your own computer vision projects. Remember, the techniques used in this Faster R-CNN PyTorch Tutorial are highly transferable, whether you are working in agriculture or another specialized field.
The future of agriculture is digital, and with the skills you’ve learned here, you are well-equipped to lead that transformation. Thank you for following this tutorial. I encourage you to take this code, adapt it to your own datasets, and build something incredible. The world of AI is constantly evolving, and by mastering these fundamental detection techniques, you are setting yourself up for success in the next generation of technological innovation.
Connect ☕ Buy me a coffee — https://ko-fi.com/eranfeit
🖥️ Email : feitgemel@gmail.com
🌐 https://eranfeit.net
🤝 Fiverr : https://www.fiverr.com/s/mB3Pbb
Enjoy,
Eran